| HOME | INSTRUMENTATION | NMR TIME | USERS & RESEARCH |
INDUSTRIAL AFFILIATES |
NEWS & LINKS |
Publications: (Details Below)
Wang, Y., Zhao, S., Somerville, R.L. and Jardetzky, O. "Solution Structure of
the DNA-binding Domain of the TyrR protein from Haemophilus Influenzae"
(2001) Protein Science 10, 592-598.
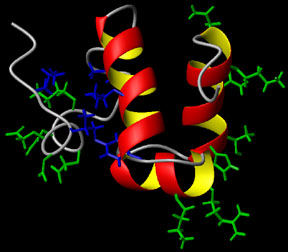
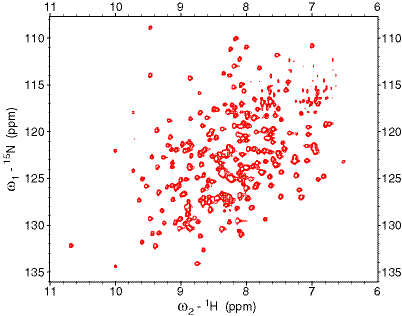
Presentations: Wang, Y. and Jardetzky, O. "Structural study of TyrR protein of H. Influenzae" Molecular Pharmacology Department Retreat, Sept 20-22, 2000, Asilomar. Details: Solution Structure of the DNA-binding Domain of the TyrR protein of Haemophilus Influenzae Wang, Y., Zhao, S., Somerville, R.L. and Jardetzky, O. (2001) Protein Science 10, 592-598. Abstract: The TyrR protein of Haemophilus influenzae is a 36-kD transcription factor whose major function is to control the expression of genes important in the biosynthesis and transport of aromatic amino acids. Using 1H and 15N NMR spectroscopy, we have determined the 3D solution structure of the TyrR C-terminal DNA-binding domain (DBD) containing residues from 258 to 318 (TyrR[258-318]). The NMR results show that this segment of TyrR consists of a potential hinge helix at its N terminus (residues 263-270) as well as three well-defined alpha-helices extending from residues 277-289 (HR-2), 293-300 (HR-1), and 304-314 (HR). Helix HR-1 and HR fold in a typical helix-turn-helix (HTH) motif. The three helices and the hinge helix are tightly bound together by hydrophobic interaction and hydrogen bonds. Several hydrophilic residues whose side chains may directly interact with DNA are identified. A hydrophobic patch that may be part of the interaction surface between the domains of TyrR protein is also observed. Comparisons with the structures of other HTH DNA-binding proteins reveal that in terms of the spatial orientation of the three helices, this protein most closely resembles the cap family.
|
| Bio-X | Structural Biology | School of Medicine | Stanford University |